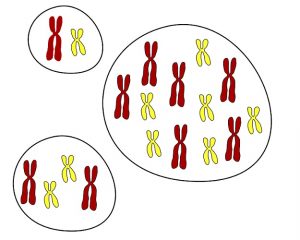
Figure 1. Haploid (n chromosomes), diploid (2n chromosomes) and hexaploid (6n chromosomes) cells. [Adapted from © Talos (CC BY-SA 3.0) via Wikimedia Commons]
Polyploidy occurs in two ways:
The same chromosomal set can be repeated several times (autopolyploidy), following unfinished mitoses in an individual.
There may be gamete fusions between two neighbouring species, resulting in individuals with the genomes of both species (allopolyploidy).
Figure 2. Soft wheat grains (left); Rye (centre) and Triticale (right). Triticale has grains of intermediate size between those of Wheat and Rye from which it is derived. [Source: © USDA] Sorghum versicolor which is diploid (2N = 10 chromosomes), S. vulgare which is tetraploid (20 chromosomes) and S. halepense which is octoploid (40 chromosomes). Alloploidy is also quite common in plants, especially in cultivated plants. This is the case for durum wheat, which is a natural allotetraploid, and for soft wheat, which is an allohexaploid. For livestock feed, breeders have created an allo-octoploid, the “Triticale”, which adds the rye genome (Secale cereale ) to soft wheat (Triticum aestivum ) (Figure B). Farmers appreciate it for its hardiness. More recently, researchers have created Tritordeum , a new species that has both barley and durum wheat genomes. Marketed since 2006, it combines the baking value of wheat with the drought tolerance of barley.
Polyploidy, under either of its two modalities, is a very effective speciation mechanism. It is estimated that more than 70% of wild species in plants are polyploids.