Symbiosis and evolution: at the origin of the eukaryotic cell
PDF
The cell of eukaryotic organisms (animals, plants, fungi) differs from that of prokaryotic organisms (Archaea and Bacteria) by the presence of several specialized organelles, such as: the nucleus (containing the genetic information of the cell), the mitochondria (site of cellular respiration), or the chloroplast (site of photosynthesis in plants). The existence and organization of mitochondrial and chloroplast DNA, as well as their biochemistry and some structural traits, have led to their being considered as ancient bacteria integrated into a host cell by an endosymbiosis process. One possible hypothesis would be that current eukaryotes would descend from an archaeal ancestor who acquired a proteobacteria, the present mitochondria. Once this step was established, some cells would have incorporated cyanobacteria that are the origin of the chloroplast. At the same time, they have acquired the ability to carry out photosynthesis, and thus an autotrophic metabolism, a particularity of plants. Throughout the process, gene transfer phenomena between symbionts, the taking over of the coding of some organelle proteins by the nucleus and the relocation of gene products into the organelles have closely integrated these prokaryotes within the host cell. The phenomenon of endosymbiosis is therefore very largely responsible for the biodiversity of eukaryotes that appeared during evolution. Thus, photosynthesis has developed in a wide variety of organisms: red and green algae, green plants through primary endosymbiosis, brown algae and many other organisms through secondary or tertiary endosymbiosis.
1. The eukaryotic cell is a chimera
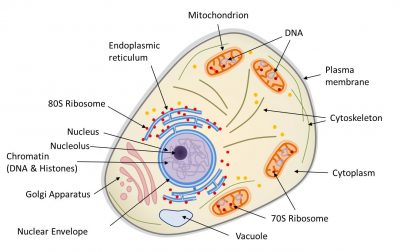
EukaryotesSingle-cell or multicellular organisms whose cells have a nucleus and organelles (endoplasmic reticulum, Golgi apparatus, various plastids, mitochondria, etc.) delimited by membranes. Eukaryotes are, along with bacteria and archaea, one of the three groups of living organisms. correspond to multicellular organisms (animals, plants, fungi) and some unicellular organisms (protozoa, for example). The main characteristic of the eukaryotic cell (Figure 1) is the existence of a nucleus (in prokaryotes, the genome is only very rarely surrounded by a membrane) surrounded by a cytoplasm containing many organelles, such as mitochondriaOrganelles of the cytoplasm of eukaryotic cells (plants, algae, animals). As the site of cellular respiration, mitochondria convert the energy of organic molecules from digestion (glucose) into energy that can be directly used by the cell (ATP) during the “Krebs cycle”. This reaction requires the presence of oxygen and releases CO2, so it plays an essential role in the carbon cycle. Mitochondria originate from a prokaryotic organism (α-proteobacteria) integrated into eukaryotic protocells 2 billion years ago. (where respiration, present in all eukaryotic cells, takes place) and chloroplastsOrganelles of the cytoplasm of photosynthetic eukaryotic cells (plants, algae). As a site of photosynthesis, chloroplasts produce O2 oxygen and play an essential role in the carbon cycle: they use light energy to fix CO2 and synthesize organic matter. They are thus responsible for the autotrophy of plants. Chloroplasts are the result of the endosymbiosis of a photosynthetic prokaryote (cyanobacterium type) in a eukaryotic cell about 1.5 billion years ago. (site for photosynthesisBioenergetic process that allows plants, algae and some bacteria to synthesize organic matter from atmospheric CO2 by using sunlight. Solar energy is used to oxidize water and reduce carbon dioxide in order to synthesize organic substances (carbohydrates). The oxidation of water leads to the formation of O2 oxygen found in the atmosphere. Photosynthesis is the basis of autotrophy, it is the result of the integrated functioning of the chloroplast within the cell., in plants in the broad sense, terrestrial plants and algae). These organelles are frequently displaced or reorganized by the cytoskeleton that triggers intracellular mobility (Figure 1).
The eukaryotic nucleus is delimited by a double membrane called the nuclear envelope (Figure 1). It contains the nuclear genome characteristic of the eukaryotic cell, i.e. the genetic material of an individual encoded in its DNA (deoxyribonucleic acid). It is usually this genome that is referred to when the genome of a eukaryote is mentioned. However, the eukaryotic cell also contains non-nuclear genomes within the organelles:
– the mitochondrial genome, within the mitochondrial matrix (Figure 1);
– the chloroplastic genome, within the chloroplast stroma (e.g. plants or algae).
The DNA constituting these three genomes is not organized in the same way. In the nucleus, the genome is distributed over several linear DNA molecules, and organized into well-differentiated chromosomes. DNA contains all coding sequences (transcribed into messenger RNA, mRNA, and translated into proteins) and non-coding sequences (not transcribed, or transcribed into RNA, but not translated). The three-dimensional configuration of the nuclear genome has a functional importance: the winding (or “condensation”) of DNA on itself and around proteins, the histonesBasic proteins associating with DNA to form the basic structure of chromatin. Histones play an important role in DNA packaging and folding, allowing a large amount of genetic information to be packaged in the tiny nucleus of a cell. Mitochondrial or chloroplastic DNA do not have the same organization: it is generally circular, rarely linear (plant mitochondria), generally without intron, and is not associated with histone proteins.
ProkaryoticMicroorganisms (usually unicellular) with a simple cellular structure, no nucleus, and almost never internal compartmentalization (the only exception being thylakoids in cyanobacteria). Two of the three groups that make up living organisms are prokaryotes: Archaea and Bacteria.-type cells (Bacteria and ArchaeaSingle-celled prokaryotic microorganisms living in particular in extreme environments (anaerobic, high salinity, very hot…). Phylogenetic research by Carl Woese and George E. Fox (1977) differentiated between archaea and other prokaryotic organisms (bacteria). Currently, living organisms are considered to consist of three groups: Archaea, Bacteria and Eukaryota.), do not have a nucleus and their DNA is circular (or -in some rare cases- linear) and organized like that of chloroplasts or mitochondria. In this way, DNA replication, transcription and translation directly take place into the cytoplasm. It should be noted, however, that Archaea are only superficially similar to Bacteria in their cellular aspect: their metabolism differs greatly, and the mechanisms and proteins involved in the replication, transcription and translation processes have similar characteristics to those of eukaryotes. Finally, prokaryotes -in general- do not have internal compartments and, if present, they are less complex (cyanobacteria are an example of an exception). Above all, compartments, when they exist, are not mobile in the cell: the cytoskeleton, which is beginning to be discovered, does not move the cellular components within it.
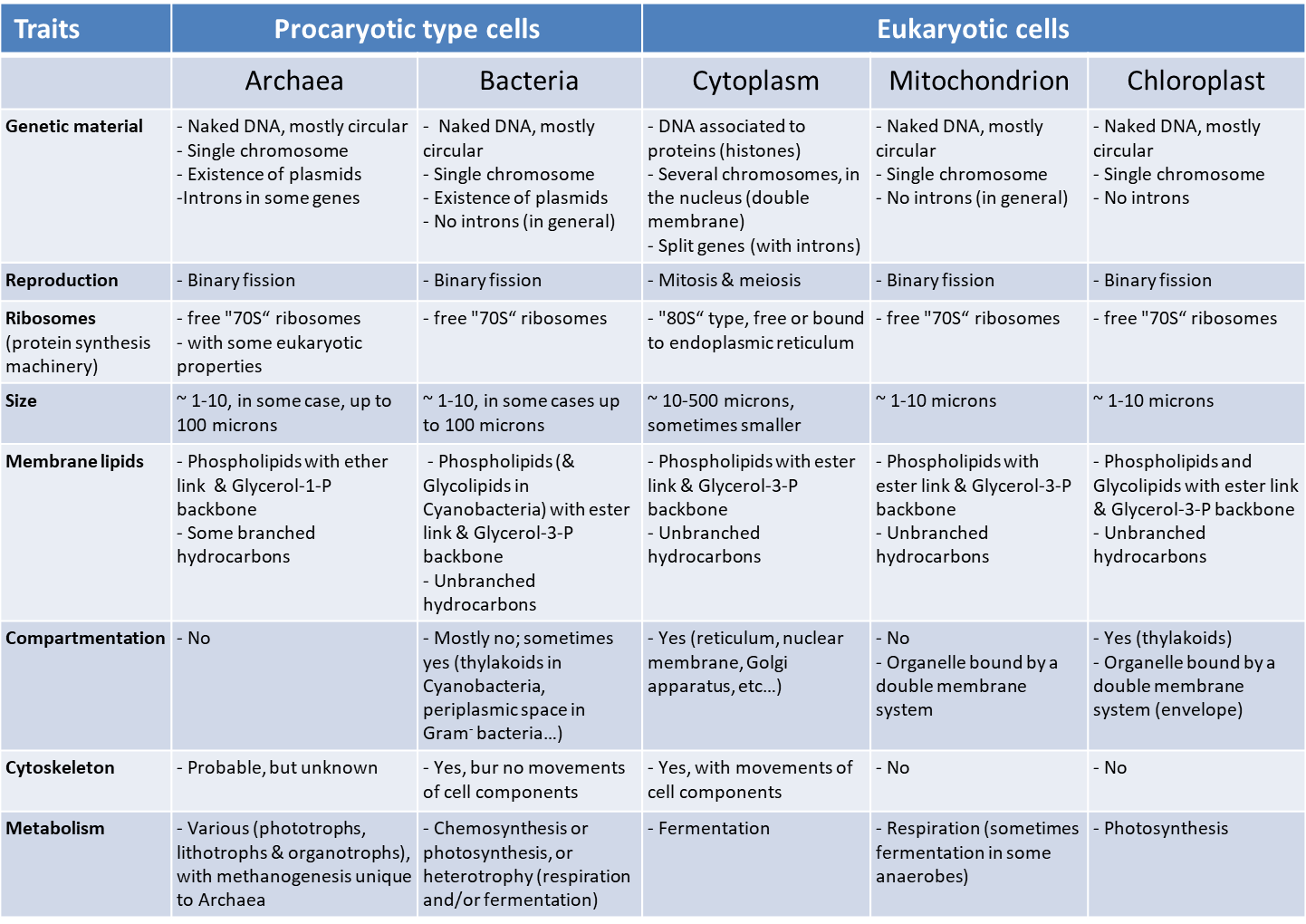
Table 1. Comparison of eukaryotic and prokaryotic cells
Table 1 compares the properties of prokaryotic and eukaryotic cells (with their mitochondria and possibly their chloroplasts). It shows that mitochondria and chloroplasts have many characteristics in common with those of prokaryotic cells. Beyond the structure of DNA, eukaryotic cell organelles are formed from pre-existing organelles, dividing by fission to multiply, like a bacteria. Similarly, they have the same protein synthesis machinery (free 70S ribosomesA huge complex composed of RNAs and ribosomal proteins that allows the translation of mRNAs into proteins. Common to all cells (prokaryotes and eukaryotes), the ribosome varies according to the organisms: 80S ribosome in eukaryotes and 70S ribosome in prokaryotes and cellular organelles (mitochondria, chloroplast). in the matrix or stroma) while in the cytoplasm of the eukaryotic cell, this machinery consists of 80S ribosomes, sometimes fixed on the membranes of the endoplasmic reticulumMembrane network of the eukaryotic cell cytoplasm, essential for cellular metabolism (lipid and protein synthesis, calcium storage). Associated with ribosomes, it is the place of synthesis of proteins secreted outside the cell and, on the other hand, proteins and lipids constituting the membranes of cellular organelles (Golgi apparatus, lysosomes, mitochondria, nucleus, ribosomes, vesicles…).. Finally, bacteria also have the metabolism of mitochondria (i.e. respiration) and, in some peculiar cases, of chloroplasts (i.e. photosynthesis). On the other hand, the eukaryotic cell is distinguished by the existence of an active protein network, the cytoskeleton, a self-organized system capable of mobility, which positions and displaces the organelles in the cell. Such a protein network is static, or even absent, in prokaryotes, and poorly developed in mitochondria and chloroplasts.
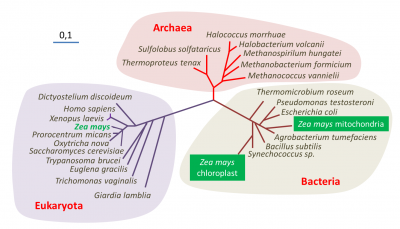
All these properties show that the eukaryotic cell is a chimera containing both characteristic components of the eukaryotic cell (the nucleus) and organelles with typically prokaryotic properties (chloroplasts, mitochondria).
The distinction between Procaryotes and Eukaryotes was proposed in 1925 by Edouard Chatton [2] (who named these two cell types), although it was not recognized until the 1950s and 1960s. The chimeric nature of eukaryotic cells had been observed since the turn of the 19th to 20th centuries. If the botanist Andreas Schimper (born in France) had the idea in 1883 that photosynthetic organisms were the result of the combination of distinct organisms, it was the Russian biologist Constantin Mereschkowsky, who first provided solid arguments that some cells come from an intracellular union of two different types of cells (endosymbiosis). In his 1905 article [3], Mereschkowsky proposed three essential ideas: (a) chloroplasts resemble cyanobacteria that very early in the evolution established a symbiosis with a heterotrophic host, (b) the host that acquired the plastids was itself the product of an earlier symbiosis between a larger, heterotrophic, amoeboid-type host cell and a smaller, micrococcus-type endosymbionte that formed the nucleus, and (c) the autotrophy of plants is entirely inherited from cyanobacteria. Mereschkowsky had not considered the origin of the mitochondria. It is to the credit of the French microbiologist Paul Portier who wrote in a text in 1918 [4] that “all living beings, all animals (…), all plants (…) are constituted by the association, the interlocking of two different beings. Each living cell contains (…) formations that cytologists refer to as “mitochondria”. For me, these organelles would be nothing more than symbiotic bacteria, what I call symbionts. “These observations received no more attention from scientists than that, and the theory fell into disgrace, especially because plastids and mitochondria were not successfully cultured, which in the 19th century was considered as evidence of a bacterial nature [5]. It took new methods of studying the cell using electron microscopy, biochemistry and molecular biology for the theory of the endosymbiotic origin of organelles in the eukaryotic cell to be brought back to life around 1970 by the American microbiologist Lynn Margulis.
2. How did the eukaryotic cell evolved?
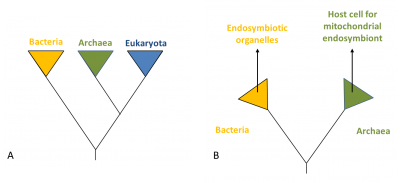
Several hypotheses have been put forward to explain the appearance of the eukaryotic cell about 1.5 billion years ago, almost a billion years after the first prokaryotic organisms appeared on Earth. This question can be addressed in very different ways, depending on whether one considers palaeontological evidence, energy aspects, the origin of the characteristics of the eukaryotic cell or the relationships of the different prokaryotic and eukaryotic lines with respect to each other [6]. Figure 3 shows hypothesis for the origin of eukaryotes and other lines (Archaea and Bacteria).
Some models assume that eukaryotes emerged from a single ancestral lineage via successive mutations during the evolutionary process. Other models postulate that eukaryotes have emerged from a symbiotic association of prokaryotic cells whose fusion would have resulted in the transition from prokaryotic to eukaryotic. These various hypotheses can be partly tested by experience, in particular through the analysis of the genomes of current organisms (prokaryotes or eukaryotes) [7,8].
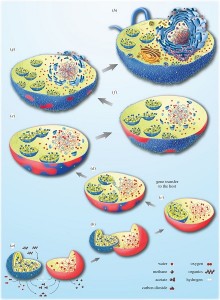
In this context, Bacteria and Archaea tend to interact closely in the same way as many current symbiotic associations. This can lead, in principle, to the situation described in Figure 4, in which the bacterial symbiote will be retained by the archeal host and eventually reside inside. In this case, the host does not feed on the symbiote, so integration is not due to a phenomenon of phagocytosisThe process by which a cell encompasses and then digests a foreign substance or organism (e g. bacteria) – although there is no doubt that phagocytosis has increased the frequency of integration of endosymbiotes in the evolution of eukaryotic cells [5].
The question of the origin of the eukaryotic cell is also linked to that of the nucleus, the emblematic structure of this cell. The establishment of a new membrane system, the nuclear membrane, in the host after mitochondria acquisition could be due to the aggregation of membrane vesicles composed of bacterial lipids. This separation between nucleus and cytoplasm could have responded to the need to separate, following gene transfer between host and symbiote, the splicing of RNA from DNA translation. It is then the selective pressure that would have led to the fixation of the compartmentalization between the newly formed nucleus and the cytoplasm [5] (Figure 4).
Thus, all eukaryotes currently known would descend from an archaean ancestor who acquired a proteobacterium during the Precambrian period, which became the mitochondria. This step is crucial: the integration of the mitochondria is therefore inseparable from the appearance of the eukaryotic cell as we know it today. The strong energy constraints exerted on the organization of prokaryotic cells were a major factor of innovation at the origin of the evolution of this cell. Only cells that had mitochondria had sufficient energy resources to reach the complexity of the eukaryotic cell, which is why there are no real intermediaries in the transition from prokaryotes to eukaryotes. It is often considered that it is only after this stage has been established that some of these cells have acquired the characteristics of the eukaryotic cell (nucleus, compartmentalization) and, eventually, integrated cyanobacteria. At the same time, they have acquired the ability to carry out photosynthesis, which is the origin of chloroplast, thus giving them an autotrophic metabolism.
Recently, some work has nuanced the age of mitochondrial symbiosis [10]. They are based on the age of acquisition of bacterial genes present in the eukaryotic nucleus (i.e. the date of their divergence from homologous genes currently found in free bacteria). They revealed that many genes, some of which contribute to the complexity of the eukaryotic cell, were most likely acquired before mitochondria. This does not imply that the cell into which the mitochondria entered was as complex as it is now, but it may have already been capable of phagocytosis, for example. This feature (specific to eukaryotes because it depends on the mobility of the cytoskeleton) could have helped in the placement of the mitochondria. The development of eukaryotic complexity therefore remains speculative, but may have begun before mitochondria, even if it undoubtedly benefited from it afterwards.
3. The endosymbiotic origin of the chloroplast
During phagocytosis in white blood cells or many protozoa (Figure 5), ingested cells are often directly digested (as in the case of prey), but sometimes they are permanently lodged in the cells (endosymbiotes). In the endosymbiosis process, organelle therefore results from internalization by phagocytosis without digestion of a prokaryote within a eukaryote (Figure 5). This is the case for chloroplasts in terrestrial plants, but also for red and green algae that are close to them [11,12].
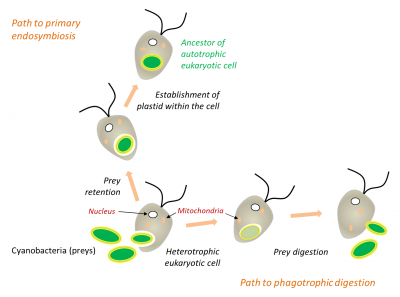
Primary and secondary endosymbioses
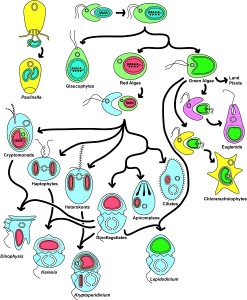
During the evolution, several endosymbiosis events repeated themselves and led to the formation of particular organisms. In primary endosymbiosis, the eukaryotic cell integrates a living prokaryotic. Thus, the chloroplasts of green line plants (red and green algae, to which terrestrial plants are attached) are derived from primary endosymbioses involving cyanobacteria. In some eukaryotes, mitochondria have evolved as a result of adaptation to anaerobic environments, but have never disappeared: they have produced particular mitochondria (hydrogenosomesOrganelles producing hydrogen, derived from a mitochondria. It is found in some anaerobic ciliates, Trichomonas, and fungi.) carrying out H2-producing fermentation (for example in some Ciliates) [14], but also small organelles, only involved in biosynthesis for the host cell, the mitosomesOrganelles present in some single-celled eukaryotic organisms, probably lacking DNA but with biosynthesis functions. [15].
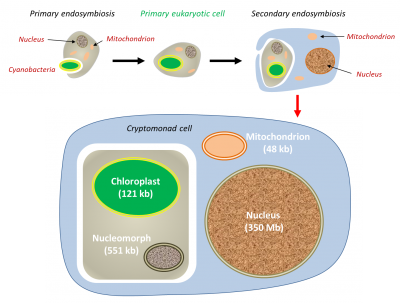
4. Integration of the prokaryote into the eukaryotic cell
All these lines have a common characteristic: a strong genetic regression of endosymbiotes. Compared to free proteobacteria such as Escherichia coli, mitochondria have lost 99% of their genes. In the extreme, hydrogenosomes and mitosomes simply no longer have a genome! Green line plastids show a 95% genetic regression compared to single-celled free cyanobacteria: the number of genes has increased from several thousand in cyanobacteria to about 100 to 200 in chloroplasts… or even none in the regressed plastids of the parasitic plant Rafflesia.
The cause of this regression is obviously the loss of genes necessary for free living, or even for certain metabolic functions. For example, as with all Gram-bacteria detected by a staining technique called Gram staining: they then appear pink under the microscope. The staining technique is based on the membrane and wall characteristics of the bacteria. However, this is not a phylogenetic classification factor: indeed, the’Gram +‘ and’Gram –‘ groups are both non-monophyletic., a layer of peptidoglycancomposed of the wall of Gram positive and Gram negative bacteria. Consists of a carbohydrate part (= polysaccharide) and a peptide part. It maintains the shape of the cells and provides mechanical protection against osmotic pressure. is located between the two membranes of cyanobacteria, essential for maintaining the structure of bacteria in the natural environment, of low osmolarityNumber of moles of “osmotically active” particles in solution in 1 litre of solution. Concept related to the osmotic pressure exerted by the particles in solution, and responsible for osmosis. Sucrose is a small osmotically active molecule while starch is a huge osmotically inactive glucose polymer. The accumulation of sucrose in a compartment leads to an increase in osmotic pressure in that compartment, which is not the case with starch.. Once integrated into the host cell, the prokaryote will find itself in a medium, the cytoplasm, whose osmolarity is very close to that of its inner medium. The peptidoglycan layer then becomes useless, and the genes responsible for the placement of the peptidoglycan layer are then lost in chloroplasts (except in glaucophytesUnicellular plankton eukaryotes living in lakes, ponds or wetlands in temperate regions. They have flagella (2 of unequal length). They have chloroplasts (called cyanelles) that are blue-green in colour, due to the presence of phycocyanins and allophycocyanins in phycobilisomes. This is a group of reduced diversity.).
Although the organelle genome is regressing, the organelle protein repertoire (the proteome), when known, remains similar to that of the free bacteria proteome: proteins operating with new functions have therefore compensated for the losses. Their coding has in fact been supported by the host’s nuclear genome: genes located in the nucleus are translated into proteins in the cytosol which are addressed to the organelle through a transit peptidePeptide sequence located at the NH2-terminal end of the newly synthesized proteins in the cytoplasm and which allows them to be addressed to the specific organelle (mitochondria, etc.) where they function. We also speak of addressing peptide.. This phenomenon of relocation of the gene product into the organelle is an absolutely essential phenomenon for the integration of the prokaryote into the host cell. The addressing machinery responsible for these transfers is a converging innovation in plastids and mitochondria. It is also an example of the new functions linked to intracellular life. These machines, which allow the import of proteins synthesized in the cytosol through the two limiting membranes of mitochondria and chloroplasts, contain a large number of proteins whose evolutionary origin is complex: proteins of both prokaryotic and eukaryotic origin, encoded in the organelle and nucleus, are found there. Together, they allow the recognition of the protein being addressed, its unfolding followed by import (the protein must be maintained in an unfolded state to cross the membranes), then the cleavage of the addressing peptide before its precise location in its functional compartment [19].
What is the origin of the genes that encode in the nucleus for functions in organelles? There are actually two (Figure 7) [16]. Sometimes, original nuclear genes have replaced organelle genes: the corresponding gene product has acquired the ability to be addressed in the organelle. This activation may have in the past led to a redundancy situation whenever a gene already encoded the same function in the organelle. From this redundancy, the organelle gene could be lost without damage (Figure 7a) [16].
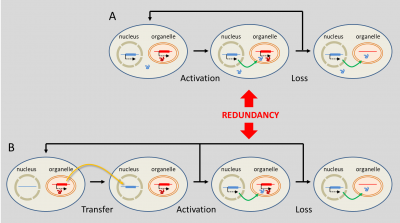
The passage of DNA fragments from organelles to the nucleus is not uncommon: large blocks of organelle DNA are inserted into the genome of some plants. These can be activated: nearly 10% of Arabidopsis thaliana‘s nuclear genes are thus derived from transfers from plastids, often followed by duplications [20]. It is not known how the DNA of the endosymbiote could have been integrated into the host genome, but it is assumed that this occurs during degradations of damaged or aged organelles accidentally releasing pieces of DNA into the host cytoplasm which are then randomly integrated into the host’s nuclear DNA.
The cytoplasmic genomes of organelles are at the crossroads of various selective forces, some of which favour their regression (such as the need for co-expression of certain genes), others favour the persistence of certain genes in the organelle genome. This could be the case for selection for a small genomic size that accelerates the multiplication of organelles and allows better transmission to daughter cells: it selects in particular the transfer of genes to the nucleus. The latter thus accumulates genetic potentialities from different lines coexisting with him in the cell [16]. Thus, while endosymbiosis reduces the genomes of endosymbiotes, it nourishes the genome of the host nucleus, contributing to its genetic diversification, and pushing for closer espouses the endosymbiotic association. Endosymbiosis therefore mixes the evolutionary lines present, by nesting but also by genetic chimerization in the nucleus of the host cell.
Finally, vertical transmission of the endosymbiote through generations is essential for the endosymbiosis to persist. Plastids must divide before the division of the host cell and must be half distributed in the two daughter cells. If their division is too rapid, they could take advantage of the host cell. On the contrary, a low division rate could lead to their disappearance. In this context, the establishment of coordination of cell division and symbiote division has been an essential element in the success of endosymbiosis. While most of the proteins involved in chloroplast division come from the cell division machinery present in cyanobacteria, some proteins are apparently of eukaryotic origin, and all are encoded in the nucleus: this is a way for the host to exercise control over chloroplast division.
5. Is symbiosis driving evolution?
More than a biological curiosity, symbiosis is certainly one of the most powerful motors of the evolution of the living world. It very quickly creates chimeric organisms that can generate new lines. It brings partners closer together and promotes massive gene transfers that also create chimeric genomes: the nuclear genome thus contains eukaryotic genes, but also genes of bacterial origin, derived from mitochondria, or even plastids, with which it borders. Such events may explain the major evolutionary leaps whose evolution seems to be punctuated, which have given rise to the great lines of life and shaped current biological diversity.
Thus, renewing the Darwinian vision of evolution by descent with modification, where one species is likely to give two, the endosymbiosis mechanisms remind us that sometimes two species, previously free and recognizable, merge into one. Man himself can be considered as an extremely integrated symbiotic community, formed by the eukaryotic cytoplasm and mitochondria, but also by the archaea and bacteria that populate, for instance, his gut microbiota…
References and notes
Cover Image : [Source Photo © Jacques Joyard]
[1] Lang T. et al (2000) Autophagy and the cvt pathway both depend on AUT9. J Bacteriol 182, 2125-2133.
[2] Chatton E. (1938) Titres et travaux scientifiques (1906-1937). Sette, Sottano, Italy. L’histoire des conditions dans lesquelles Chatton a établi le concept de procaryote et eucaryote est décrite par Sapp J. (2005) The Prokaryote-Eukaryote Dichotomy: Meanings and Mythology, Microbiol Mol Biol Rev. 69, 292–305.
[3] Mereschkowsky C. 1905 Uber Natur und Ursprung der Chromatophoren im Pflanzenreiche. Centralbl. 25, 593-604; translated by Martin W, Kowallik K. (1999) Annotated English translation of Mereschkowsky’s 1905 paper ‘Uber Natur und Ursprung der Chromatophoren im Pflanzenreiche‘. Eur. J. Phycol. 34, 287–295.
[4] Portier P. (1918) Les Symbiotes. Masson (ed.), Paris. (in french)
[5] Martin W.F., Garg S. & Zimorski V. (2015) Endosymbiotic theories for eukaryote origin. Phil. Trans. R. Soc. B370, 20140330.
[6] Selosse M.A. (2012). Gloire et disgrâce de la théorie endosymbiotique. La Recherche 468: 92-94. (in french)
[7] Archibald J.M. (2014) One plus one equals one: symbiosis and the evolution of complex life. Oxford, UK: Oxford University Press.
[8] McFadden G.I. (2014) Origin and Evolution of Plastids and Photosynthesis in Eukaryotes, Cold Spring Harb.Perspect. Biol. 6, a016105
[9] Martin W. & Müller M. (1998) The hydrogen hypothesis for the first eukaryote. Nature 392, 37-41.
[10] Ettema T.J.G. (2016) Mitochondria in the second act. Nature 531, 39-40doi:10.1038/nature16876
[11] Archibald J.M. & Keeling P.J. (2002) Recycled plastids: a “green movement” in eukaryotic evolution. Trends Genetics 18, 577-584.
[12] Keeling P.J. (2004) Diversity and evolutionary history of plastids and their hosts. Am. J. Bot. 91, 1481-1493.
[13] Douce R., Block M.A., Dorne A.J., Joyard J. (1984) The plastid envelope membranes: their structure, composition, and role in chloroplast biogenesis. Subcell. Biochem. 10, 1-84, Springer US (Ed.)
[14] Selosse M.A. & Loiseaux-de Goër S. (1997) La Saga de l’endosymbiose, La Recherche 296, 36 (in french)
[15] Embley T.M. & Martin W. (2006) Eukaryotic evolution, changes and challenges. Nature 440, 623-630
[16] Lefèvre T., Renaud F., Selosse M.-A. & Thomas F. (2010). Évolution des interactions entre espèces, in F. Thomas, T. Lefèvre & M. Raymond (ed.), Biologie évolutive, p. 530-613. De Boeck, Paris. (in french)
[17] Keeling P.J. (2010) The endosymbiotic origin, diversification and fate of plastids. Phil. Trans. R. Soc. B 365, 729-748
[18] Douglas S. et al (2001) The highly reduced genome of an enslaved algal nucleus. Nature 410, 1091-1096.
[19] Selosse M.A., Albert B. & Godelle B. (2001) Small is successful: selection for reducing organelle’s genome size favours gene transfer to the nucleus. Trends Ecol Evol 16, 135-141.
[20] Jarvis P. (2004) Organellar Proteomics: Chloroplasts in the Spotlight. Current Biology 14, R317-9. http://www.cell.com/current-biology/references/S0960-9822%2804%2900231-3
The Encyclopedia of the Environment by the Association des Encyclopédies de l'Environnement et de l'Énergie (www.a3e.fr), contractually linked to the University of Grenoble Alpes and Grenoble INP, and sponsored by the French Academy of Sciences.
To cite this article: SELOSSE Marc-André, JOYARD Jacques (May 6, 2026), Symbiosis and evolution: at the origin of the eukaryotic cell, Encyclopedia of the Environment, Accessed May 29, 2026 [online ISSN 2555-0950] url : https://www.encyclopedie-environnement.org/en/life/symbiosis-and-evolution-origin-eukaryotic-cell-2/.
The articles in the Encyclopedia of the Environment are made available under the terms of the Creative Commons BY-NC-SA license, which authorizes reproduction subject to: citing the source, not making commercial use of them, sharing identical initial conditions, reproducing at each reuse or distribution the mention of this Creative Commons BY-NC-SA license.